publications
publications by categories in reversed chronological order. generated by jekyll-scholar.
2026
- ICLR
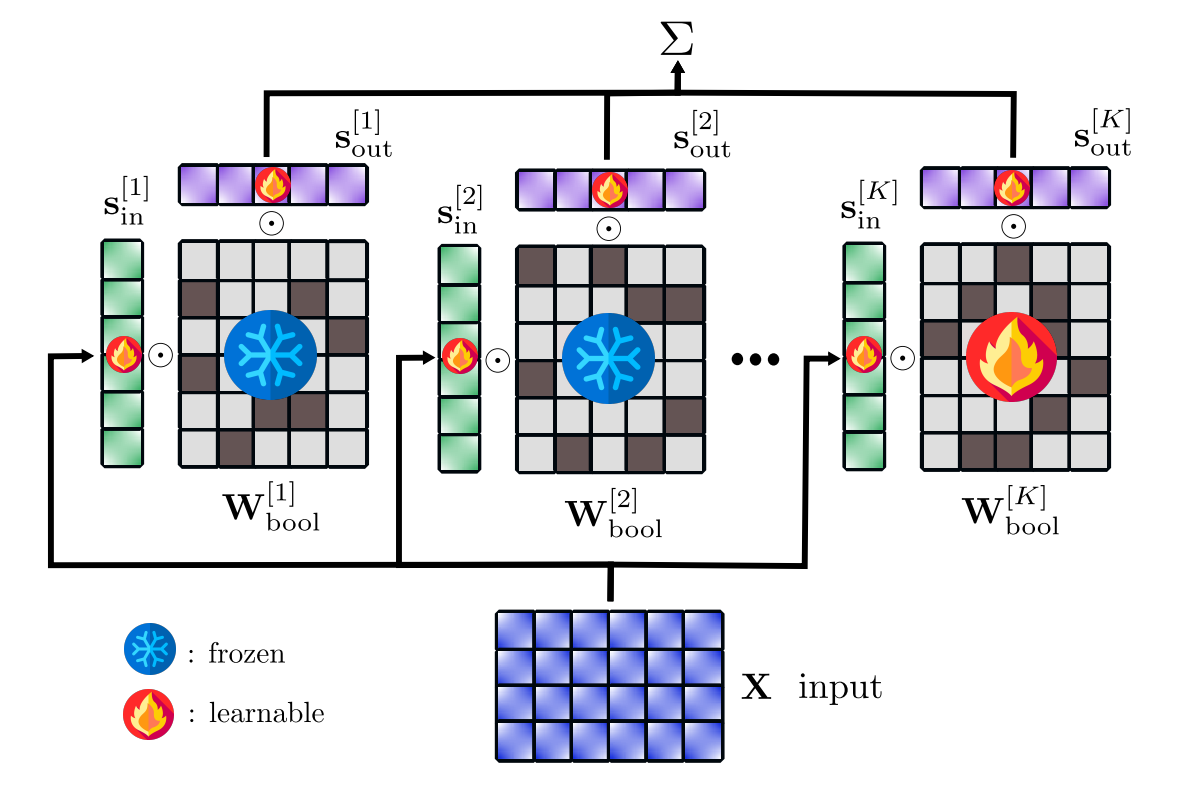 Highly Efficient and Effective LLMs with Multi-Boolean ArchitecturesBa-Hien Tran and Van Minh NguyenIn The Fourteenth International Conference on Learning Representations, 2026
Highly Efficient and Effective LLMs with Multi-Boolean ArchitecturesBa-Hien Tran and Van Minh NguyenIn The Fourteenth International Conference on Learning Representations, 2026Weight binarization has emerged as a promising strategy to reduce the complexity of large language models (LLMs). Existing approaches fall into post-training binarization, which is simple but causes severe performance loss, and training-aware methods, which depend on full-precision latent weights, adding complexity and limiting efficiency. We propose a novel framework that represents LLMs with multi-kernel Boolean parameters and, for the first time, enables direct finetuning LMMs in the Boolean domain, eliminating the need for latent weights. This enhances representational capacity and dramatically reduces complexity during both finetuning and inference. Extensive experiments across diverse LLMs show our method outperforms recent ultra low-bit quantization and binarization techniques.
@inproceedings{tran2026highly, title = {Highly Efficient and Effective {LLM}s with Multi-Boolean Architectures}, author = {Tran, Ba-Hien and Nguyen, Van Minh}, booktitle = {The Fourteenth International Conference on Learning Representations}, year = {2026}, url = {https://openreview.net/forum?id=r0CH5dF3Se}, } - ICLR
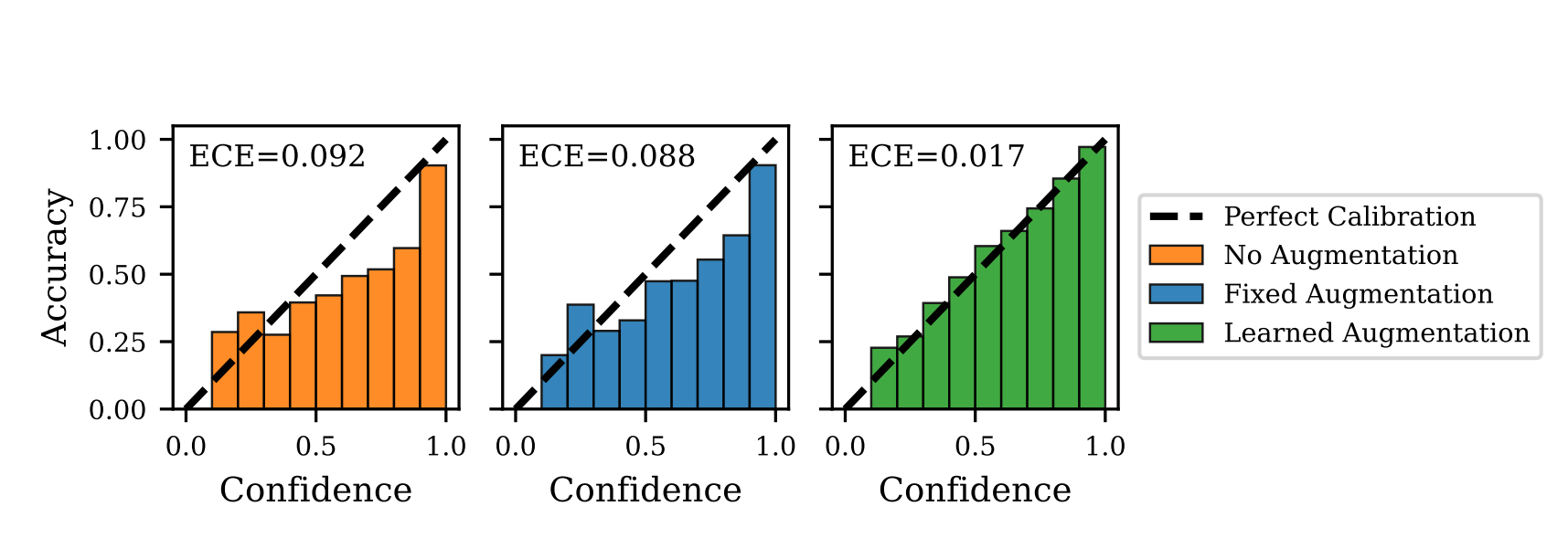 Optimizing Data Augmentation through Bayesian Model SelectionMadi Matymov, Ba-Hien Tran, Michael Kampffmeyer, and 2 more authorsIn The Fourteenth International Conference on Learning Representations, 2026
Optimizing Data Augmentation through Bayesian Model SelectionMadi Matymov, Ba-Hien Tran, Michael Kampffmeyer, and 2 more authorsIn The Fourteenth International Conference on Learning Representations, 2026Data Augmentation (DA) has become an essential tool to improve robustness and generalization of modern machine learning. However, when deciding on DA strategies it is critical to choose parameters carefully, and this can be a daunting task which is traditionally left to trial-and-error or expensive optimization based on validation performance. In this paper, we counter these limitations by proposing a novel framework for optimizing DA. In particular, we take a probabilistic view of DA, which leads to the interpretation of augmentation parameters as model (hyper)-parameters, and the optimization of the marginal likelihood with respect to these parameters as a Bayesian model selection problem. Due to its intractability, we derive a tractable ELBO, which allows us to optimize augmentation parameters jointly with model parameters. We provide extensive theoretical results on variational approximation quality, generalization guarantees, invariance properties, and connections to empirical Bayes. Through experiments on computer vision and NLP tasks, we show that our approach improves calibration and yields robust performance over fixed or no augmentation. Our work provides a rigorous foundation for optimizing DA through Bayesian principles with significant potential for robust machine learning.
@inproceedings{matymov2026optimizing, title = {Optimizing Data Augmentation through Bayesian Model Selection}, author = {Matymov, Madi and Tran, Ba-Hien and Kampffmeyer, Michael and Heinonen, Markus and Filippone, Maurizio}, booktitle = {The Fourteenth International Conference on Learning Representations}, year = {2026}, url = {https://openreview.net/forum?id=ofYuPZ0sK0}, }
2025
- AISTATS
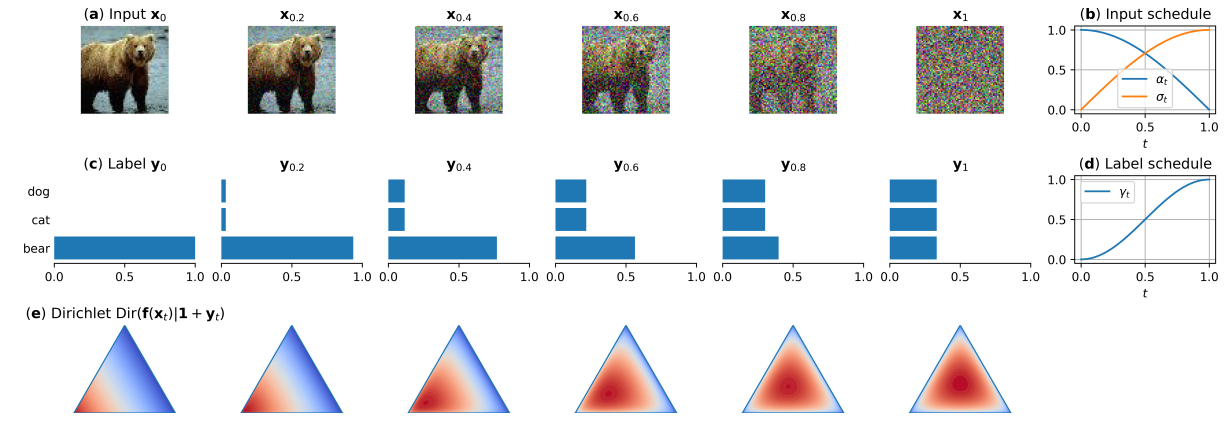 Robust Classification by Coupling Data Mollification with Label SmoothingMarkus Heinonen, Ba-Hien Tran, Michael Kampffmeyer, and 1 more authorIn Proceedings of The 28th International Conference on Artificial Intelligence and Statistics, 2025
Robust Classification by Coupling Data Mollification with Label SmoothingMarkus Heinonen, Ba-Hien Tran, Michael Kampffmeyer, and 1 more authorIn Proceedings of The 28th International Conference on Artificial Intelligence and Statistics, 2025Introducing training-time augmentations is a key technique to enhance generalization and prepare deep neural networks against test-time corruptions. Inspired by the success of generative diffusion models, we propose a novel approach of coupling data mollification, in the form of image noising and blurring, with label smoothing to align predicted label confidences with image degradation. The method is simple to implement, introduces negligible overheads, and can be combined with existing augmentations. We demonstrate improved robustness and uncertainty quantification on the corrupted image benchmarks of CIFAR, TinyImageNet and ImageNet datasets.
@inproceedings{pmlr-v258-heinonen25a, title = {Robust Classification by Coupling Data Mollification with Label Smoothing}, author = {Heinonen, Markus and Tran, Ba-Hien and Kampffmeyer, Michael and Filippone, Maurizio}, booktitle = {Proceedings of The 28th International Conference on Artificial Intelligence and Statistics}, pages = {4960--4968}, year = {2025}, editor = {Li, Yingzhen and Mandt, Stephan and Agrawal, Shipra and Khan, Emtiyaz}, volume = {258}, series = {Proceedings of Machine Learning Research}, publisher = {PMLR}, url = {https://proceedings.mlr.press/v258/heinonen25a.html}, } - Preprint
 Universal Adaptive Environment DiscoveryMadi Matymov, Ba-Hien Tran, and Maurizio FilipponearXiv preprint arXiv:2510.12547, 2025
Universal Adaptive Environment DiscoveryMadi Matymov, Ba-Hien Tran, and Maurizio FilipponearXiv preprint arXiv:2510.12547, 2025An open problem in Machine Learning is how to avoid models to exploit spurious correlations in the data; a famous example is the background-label shortcut in the Waterbirds dataset. A common remedy is to train a model across multiple environments; in the Waterbirds dataset, this corresponds to training by randomizing the background. However, selecting the right environments is a challenging problem, given that these are rarely known a priori. We propose Universal Adaptive Environment Discovery (UAED), a unified framework that learns a distribution over data transformations that instantiate environments, and optimizes any robust objective averaged over this learned distribution. UAED yields adaptive variants of IRM, REx, GroupDRO, and CORAL without predefined groups or manual environment design. We provide a theoretical analysis by providing PAC-Bayes bounds and by showing robustness to test environment distributions under standard conditions. Empirically, UAED discovers interpretable environment distributions and improves worst-case accuracy on standard benchmarks, while remaining competitive on mean accuracy. Our results indicate that making environments adaptive is a practical route to out-of-distribution generalization.
@article{matymov2025universal, title = {Universal Adaptive Environment Discovery}, author = {Matymov, Madi and Tran, Ba-Hien and Filippone, Maurizio}, journal = {arXiv preprint arXiv:2510.12547}, year = {2025}, }
2024
- NeurIPS
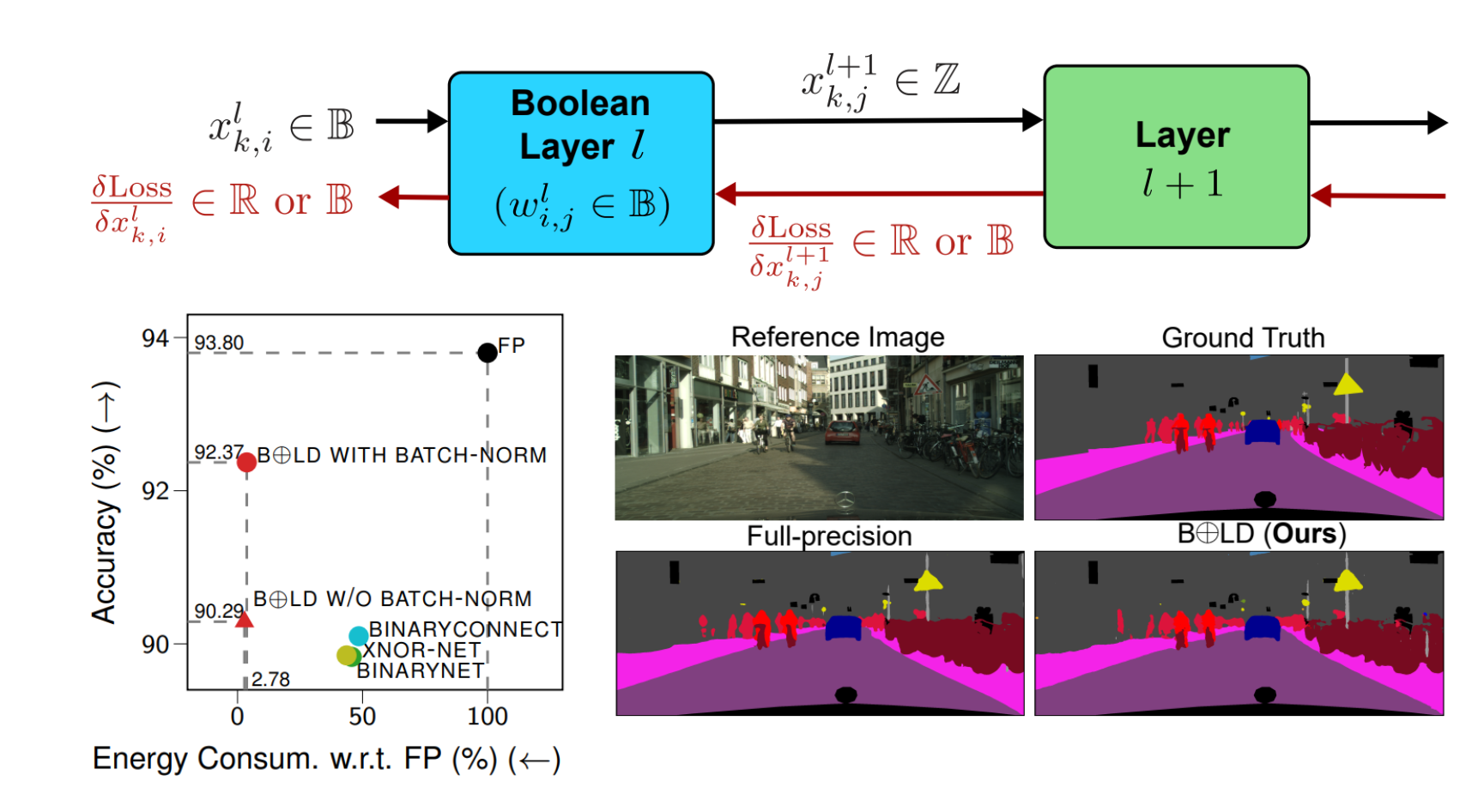 BOLD: Boolean Logic Deep LearningVan Minh Nguyen, Cristian Ocampo, Aymen Askri, and 2 more authorsIn Advances in Neural Information Processing Systems, 2024
BOLD: Boolean Logic Deep LearningVan Minh Nguyen, Cristian Ocampo, Aymen Askri, and 2 more authorsIn Advances in Neural Information Processing Systems, 2024Computational intensiveness of deep learning has motivated low-precision arithmetic designs. However, the current quantized/binarized training approaches are limited by: (1) significant performance loss due to arbitrary approximations of the latent weight gradient through its discretization/binarization function, and (2) training computational intensiveness due to the reliance on full-precision latent weights. This paper proposes a novel mathematical principle by introducing the notion of Boolean variation such that neurons made of Boolean weights and/or activations can be trained —for the first time— natively in Boolean domain instead of latent-weight gradient descent and real arithmetic. We explore its convergence, conduct extensively experimental benchmarking, and provide consistent complexity evaluation by considering chip architecture, memory hierarchy, dataflow, and arithmetic precision. Our approach achieves baseline full-precision accuracy in ImageNet classification and surpasses state-of-the-art results in semantic segmentation, with notable performance in image super-resolution, and natural language understanding with transformer-based models. Moreover, it significantly reduces energy consumption during both training and inference.
@inproceedings{nguyen2024bold, author = {Nguyen, Van Minh and Ocampo, Cristian and Askri, Aymen and Leconte, Louis and Tran, Ba-Hien}, booktitle = {Advances in Neural Information Processing Systems}, editor = {Globerson, A. and Mackey, L. and Belgrave, D. and Fan, A. and Paquet, U. and Tomczak, J. and Zhang, C.}, pages = {61912--61962}, publisher = {Curran Associates, Inc.}, title = {BOLD: Boolean Logic Deep Learning}, url = {https://proceedings.neurips.cc/paper_files/paper/2024/file/718a3c5cf135894db6e718725f52ef9a-Paper-Conference.pdf}, volume = {37}, year = {2024}, } - SpatialStats
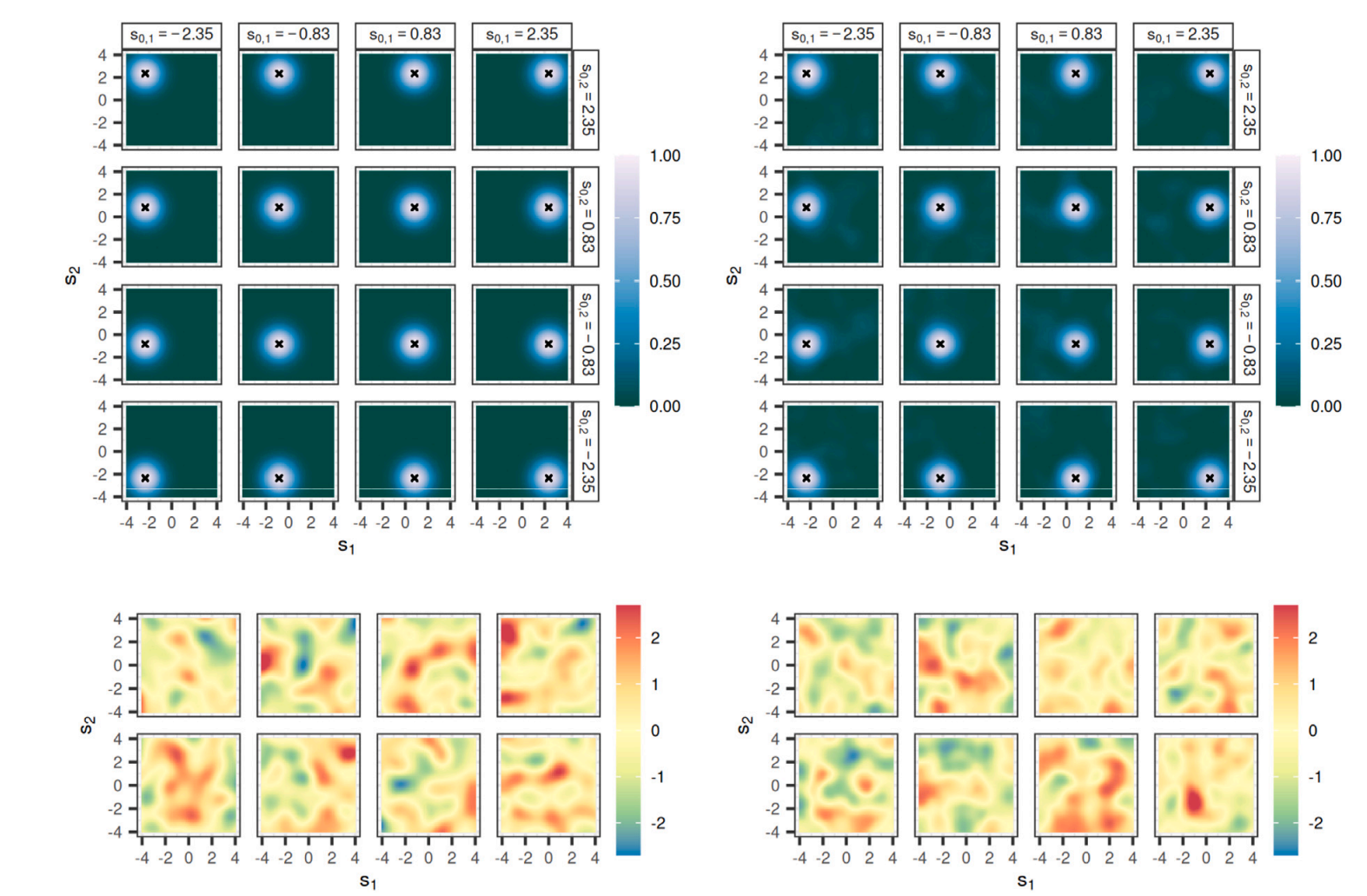 Spatial Bayesian Neural NetworksAndrew Zammit-Mangion, Michael D Kaminski, Ba-Hien Tran, and 2 more authorsSpatial Statistics, 2024
Spatial Bayesian Neural NetworksAndrew Zammit-Mangion, Michael D Kaminski, Ba-Hien Tran, and 2 more authorsSpatial Statistics, 2024Statistical models for spatial processes play a central role in analyses of spatial data. Yet, it is the simple, interpretable, and well understood models that are routinely employed even though, as is revealed through prior and posterior predictive checks, these can poorly characterise the spatial heterogeneity in the underlying process of interest. Here, we propose a new, flexible class of spatial-process models, which we refer to as spatial Bayesian neural networks (SBNNs). An SBNN leverages the representational capacity of a Bayesian neural network; it is tailored to a spatial setting by incorporating a spatial “embedding layer” into the network and, possibly, spatially-varying network parameters. An SBNN is calibrated by matching its finite-dimensional distribution at locations on a fine gridding of space to that of a target process of interest. That process could be easy to simulate from or we may have many realisations from it. We propose several variants of SBNNs, most of which are able to match the finite-dimensional distribution of the target process at the selected grid better than conventional BNNs of similar complexity. We also show that an SBNN can be used to represent a variety of spatial processes often used in practice, such as Gaussian processes, lognormal processes, and max-stable processes. We briefly discuss the tools that could be used to make inference with SBNNs, and we conclude with a discussion of their advantages and limitations.
@article{zammit2024spatial, title = {Spatial Bayesian Neural Networks}, author = {Zammit-Mangion, Andrew and Kaminski, Michael D and Tran, Ba-Hien and Filippone, Maurizio and Cressie, Noel}, journal = {Spatial Statistics}, volume = {60}, pages = {100825}, year = {2024}, publisher = {Elsevier}, }
2023
- ICML
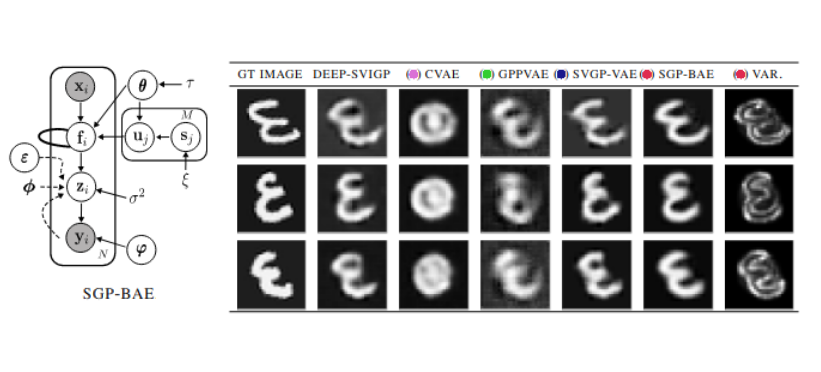 Fully Bayesian Autoencoders with Latent Sparse Gaussian ProcessesBa-Hien Tran, Babak Shahbaba, Stephan Mandt, and 1 more authorIn Proceedings of the 40th International Conference on Machine Learning, 2023
Fully Bayesian Autoencoders with Latent Sparse Gaussian ProcessesBa-Hien Tran, Babak Shahbaba, Stephan Mandt, and 1 more authorIn Proceedings of the 40th International Conference on Machine Learning, 2023We present a fully Bayesian autoencoder model that treats both local latent variables and global decoder parameters in a Bayesian fashion. This approach allows for flexible priors and posterior approximations while keeping the inference costs low. To achieve this, we introduce an amortized MCMC approach by utilizing an implicit stochastic network to learn sampling from the posterior over local latent variables. Furthermore, we extend the model by incorporating a Sparse Gaussian Process prior over the latent space, allowing for a fully Bayesian treatment of inducing points and kernel hyperparameters and leading to improved scalability. Additionally, we enable Deep Gaussian Process priors on the latent space and the handling of missing data. We evaluate our model on a range of experiments focusing on dynamic representation learning and generative modeling, demonstrating the strong performance of our approach in comparison to existing methods that combine Gaussian Processes and autoencoders.
@inproceedings{tran2023fully, title = {Fully {B}ayesian Autoencoders with Latent Sparse {G}aussian Processes}, author = {Tran, Ba-Hien and Shahbaba, Babak and Mandt, Stephan and Filippone, Maurizio}, booktitle = {Proceedings of the 40th International Conference on Machine Learning}, pages = {34409--34430}, year = {2023}, editor = {Krause, Andreas and Brunskill, Emma and Cho, Kyunghyun and Engelhardt, Barbara and Sabato, Sivan and Scarlett, Jonathan}, volume = {202}, series = {Proceedings of Machine Learning Research}, publisher = {PMLR}, url = {https://proceedings.mlr.press/v202/tran23a.html}, } - PhD Thesis
 Advancing Bayesian Deep Learning : Sensible Priors and Accelerated InferenceBa-Hien TranOct 2023
Advancing Bayesian Deep Learning : Sensible Priors and Accelerated InferenceBa-Hien TranOct 2023Over the past decade, deep learning has witnessed remarkable success in a wide range of applications, revolutionizing various fields with its unprecedented performance. However, a fundamental limitation of deep learning models lies in their inability to accurately quantify prediction uncertainty, posing challenges for applications that demand robust risk assessment. Fortunately, Bayesian deep learning provides a promising solution by adopting a Bayesian formulation for neural networks. Despite significant progress in recent years, there remain several challenges that hinder the widespread adoption and applicability of Bayesian deep learning. In this thesis, we address some of these challenges by proposing solutions to choose sensible priors and accelerate inference for Bayesian deep learning models. The first contribution of the thesis is a study of the pathologies associated with poor choices of priors for Bayesian neural networks for supervised learning tasks and a proposal to tackle this problem in a practical and effective way. Specifically, our approach involves reasoning in terms of functional priors, which are more easily elicited, and adjusting the priors of neural network parameters to align with these functional priors. The second contribution is a novel framework for conducting model selection for Bayesian autoencoders for unsupervised tasks, such as representation learning and generative modeling. To this end, we reason about the marginal likelihood of these models in terms of functional priors and propose a fully sample-based approach for its optimization. The third contribution is a novel fully Bayesian autoencoder model that treats both local latent variables and the global decoder in a Bayesian fashion. We propose an efficient amortized MCMC scheme for this model and impose sparse Gaussian process priors over the latent space to capture correlations between latent encodings. The last contribution is a simple yet effective approach to improve likelihood-based generative models through data mollification. This accelerates inference for these models by allowing accurate density-esimation in low-density regions while addressing manifold overfitting.
@phdthesis{tran:tel-04565603, title = {{Advancing Bayesian Deep Learning : Sensible Priors and Accelerated Inference}}, author = {Tran, Ba-Hien}, url = {https://theses.hal.science/tel-04565603}, number = {2023SORUS280}, school = {{Sorbonne Universit{\'e}}}, year = {2023}, month = oct, keywords = {Bayesian inference ; Neural networks ; Gaussian processes ; Inf{\'e}rence bay{\'e}sienne ; R{\'e}seau neuronal ; Processus gaussiens}, type = {Theses}, hal_id = {tel-04565603}, hal_version = {v1}, } - NeurIPS
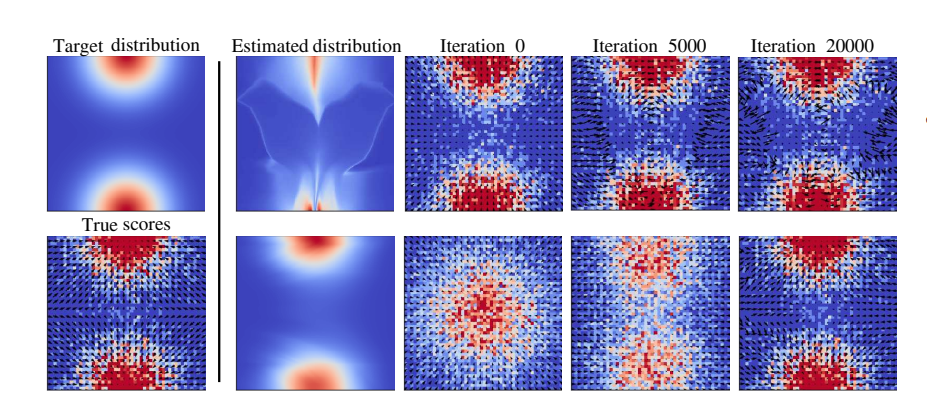 One-Line-of-Code Data Mollification Improves Optimization of Likelihood-based Generative ModelsBa-Hien Tran, Giulio Franzese, Pietro Michiardi, and 1 more authorIn Advances in Neural Information Processing Systems, 2023
One-Line-of-Code Data Mollification Improves Optimization of Likelihood-based Generative ModelsBa-Hien Tran, Giulio Franzese, Pietro Michiardi, and 1 more authorIn Advances in Neural Information Processing Systems, 2023Generative Models (GMs) have attracted considerable attention due to their tremendous success in various domains, such as computer vision where they are capable to generate impressive realistic-looking images. Likelihood-based GMs are attractive due to the possibility to generate new data by a single model evaluation. However, they typically achieve lower sample quality compared to state-of-the-art score-based Diffusion Models (DMs). This paper provides a significant step in the direction of addressing this limitation. The idea is to borrow one of the strengths of score-based DMs, which is the ability to perform accurate density estimation in low-density regions and to address manifold overfitting by means of data mollification. We propose a view of data mollification within likelihood-based GMs as a continuation method, whereby the optimization objective smoothly transitions from simple-to-optimize to the original target. Crucially, data mollification can be implemented by adding one line of code in the optimization loop, and we demonstrate that this provides a boost in generation quality of likelihood-based GMs, without computational overheads. We report results on real-world image data sets and UCI benchmarks with popular likelihood-based GMs, including variants of variational autoencoders and normalizing flows, showing large improvements in FID score and density estimation.
@inproceedings{tran2023one, author = {Tran, Ba-Hien and Franzese, Giulio and Michiardi, Pietro and Filippone, Maurizio}, booktitle = {Advances in Neural Information Processing Systems}, editor = {Oh, A. and Naumann, T. and Globerson, A. and Saenko, K. and Hardt, M. and Levine, S.}, pages = {6545--6567}, publisher = {Curran Associates, Inc.}, title = {One-Line-of-Code Data Mollification Improves Optimization of Likelihood-based Generative Models}, url = {https://proceedings.neurips.cc/paper_files/paper/2023/file/1516a7f7507d5550db5c7f29e995ec8c-Paper-Conference.pdf}, volume = {36}, year = {2023}, }
2022
- JMLR
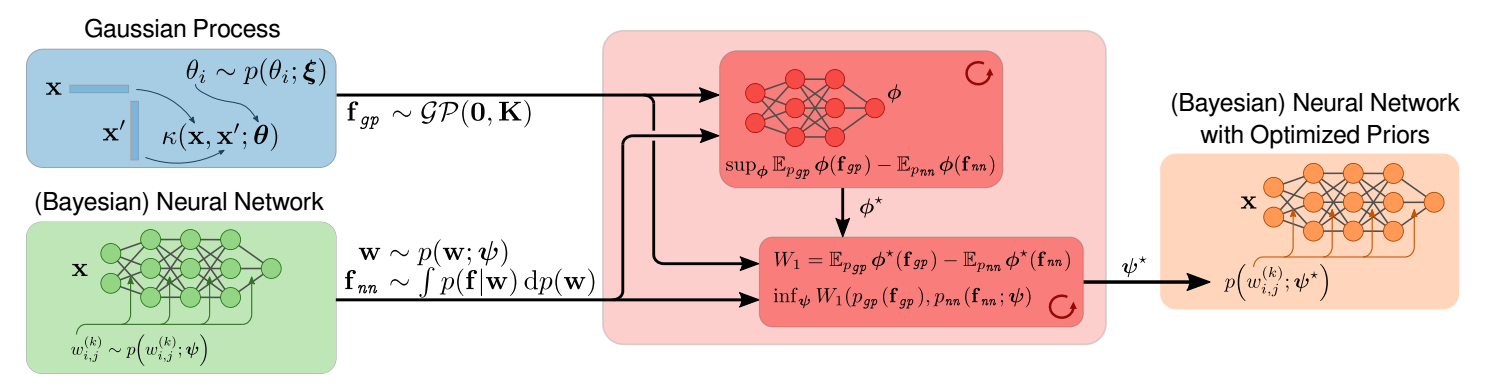 All You Need is a Good Functional Prior for Bayesian Deep LearningBa-Hien Tran, Simone Rossi, Dimitrios Milios, and 1 more authorJournal of Machine Learning Research, 2022
All You Need is a Good Functional Prior for Bayesian Deep LearningBa-Hien Tran, Simone Rossi, Dimitrios Milios, and 1 more authorJournal of Machine Learning Research, 2022The Bayesian treatment of neural networks dictates that a prior distribution is specified over their weight and bias parameters. This poses a challenge because modern neural networks are characterized by a large number of parameters, and the choice of these priors has an uncontrolled effect on the induced functional prior, which is the distribution of the functions obtained by sampling the parameters from their prior distribution. We argue that this is a hugely limiting aspect of Bayesian deep learning, and this work tackles this limitation in a practical and effective way. Our proposal is to reason in terms of functional priors, which are easier to elicit, and to “tune” the priors of neural network parameters in a way that they reflect such functional priors. Gaussian processes offer a rigorous framework to define prior distributions over functions, and we propose a novel and robust framework to match their prior with the functional prior of neural networks based on the minimization of their Wasserstein distance. We provide vast experimental evidence that coupling these priors with scalable Markov chain Monte Carlo sampling offers systematically large performance improvements over alternative choices of priors and state-of-the-art approximate Bayesian deep learning approaches. We consider this work a considerable step in the direction of making the long-standing challenge of carrying out a fully Bayesian treatment of neural networks, including convolutional neural networks, a concrete possibility.
@article{tran2022all, author = {Tran, Ba-Hien and Rossi, Simone and Milios, Dimitrios and Filippone, Maurizio}, title = {All You Need is a Good Functional Prior for Bayesian Deep Learning}, journal = {Journal of Machine Learning Research}, year = {2022}, volume = {23}, number = {74}, pages = {1--56}, url = {http://jmlr.org/papers/v23/20-1340.html} }
2021
- NeurIPS
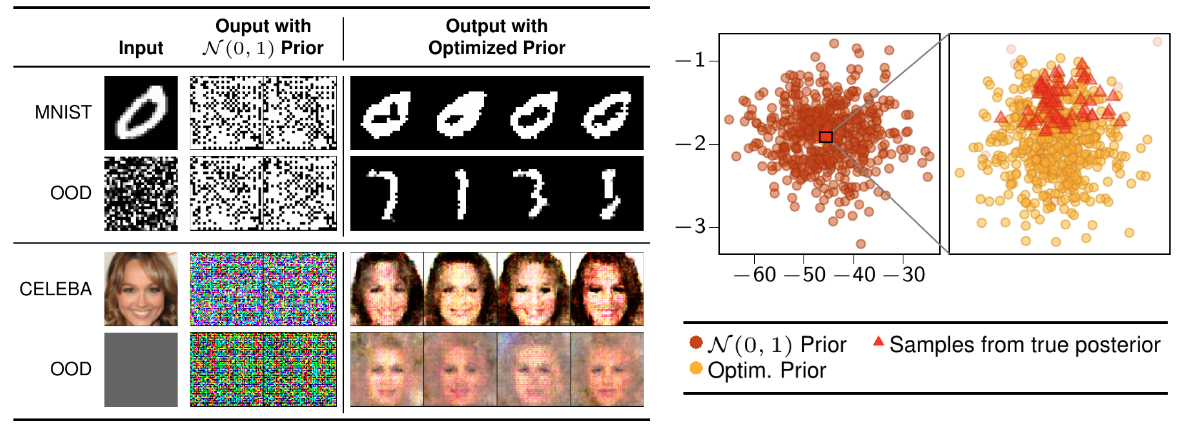 Model Selection for Bayesian AutoencodersBa-Hien Tran, Simone Rossi, Dimitrios Milios, and 3 more authorsIn Advances in Neural Information Processing Systems, 2021
Model Selection for Bayesian AutoencodersBa-Hien Tran, Simone Rossi, Dimitrios Milios, and 3 more authorsIn Advances in Neural Information Processing Systems, 2021We develop a novel method for carrying out model selection for Bayesian autoencoders (BAEs) by means of prior hyper-parameter optimization. Inspired by the common practice of type-II maximum likelihood optimization and its equivalence to Kullback-Leibler divergence minimization, we propose to optimize the distributional sliced-Wasserstein distance (DSWD) between the output of the autoencoder and the empirical data distribution. The advantages of this formulation are that we can estimate the DSWD based on samples and handle high-dimensional problems. We carry out posterior estimation of the BAE parameters via stochastic gradient Hamiltonian Monte Carlo and turn our BAE into a generative model by fitting a flexible Dirichlet mixture model in the latent space. Consequently, we obtain a powerful alternative to variational autoencoders, which are the preferred choice in modern applications of autoencoders for representation learning with uncertainty. We evaluate our approach qualitatively and quantitatively using a vast experimental campaign on a number of unsupervised learning tasks and show that, in smalldata regimes where priors matter, our approach provides state-of-the-art results, outperforming multiple competitive baselines.
@inproceedings{tran2021model, author = {Tran, Ba-Hien and Rossi, Simone and Milios, Dimitrios and Michiardi, Pietro and Bonilla, Edwin V and Filippone, Maurizio}, booktitle = {Advances in Neural Information Processing Systems}, editor = {Ranzato, M. and Beygelzimer, A. and Dauphin, Y. and Liang, P.S. and Vaughan, J. Wortman}, pages = {19730--19742}, publisher = {Curran Associates, Inc.}, title = {Model Selection for Bayesian Autoencoders}, url = {https://proceedings.neurips.cc/paper_files/paper/2021/file/a41db61e2728ef963614a8c8755b9b9a-Paper.pdf}, volume = {34}, year = {2021}, }